Covid-19 symptoms simulation (NetLogo)
Name: COVID-19 symptoms simulation
Author: Kamila Steinbartová, stek15, Jan 2021
Used tool: NetLogo 6.1.1
Contents
1. Problem definition
We are all familiar with the present pandemic situation. One of the main factors of the financial impact of the disease is the need for testing. Be it individual or nation-wide, the time, resources, and personnel required are far beyond what could be manageable long-term. Securely identifying the symptoms and tracking them could allow for more accurate quarantine measures.
The available information on the COVID-19 disease is being constantly updated and developed. To take this into consideration, the model was created to be adaptable and easily editable. Even during the course of model creation, which only took a couple of weeks, the most commonly listed symptoms have been updated based on newly available information several times. The final list of 7 of the most common symptoms among patients hospitalized with COVID-19 as stated by Chinese researchers has been taken from WebMD (2021).
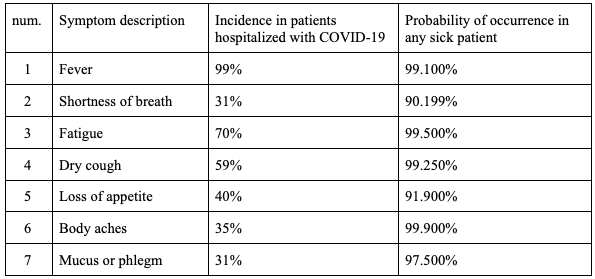
The table below lists said symptoms, their percentual incidence in patients hospitalized with COVID-19, and the added probability P(A∪B∪C) of occurrence in any sick patient (considering COVID-19, the common cold, and the flu), calculated from data taken from Health Service Executive, Ireland (2021).
The use of these parameters in my simulation is explained in the following sections.
2. Method
The tool used for this simulation is NetLogo 6.1.1.
3. Model
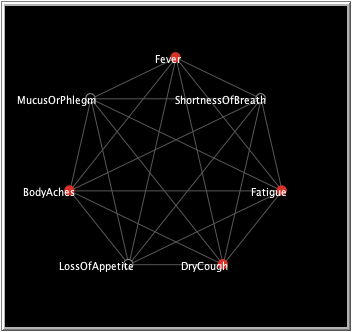
This model is a representation of the COVID-19 virus, or rather of the effects said virus has on an affected person. Each node of the model represents one of the 7 symptoms listed above. The model simulates the development of the symptoms in a patient. If the symptom turns red, it became activated (experienced). Upon deactivation, it turns black again. Each symptom is connected to all the others in order to create a conclusive network.
Before the simulation is started, the user is to choose the initial settings for the model using the two sliders on the left side of the model: contraction-susceptibility and exposure-to-virus. With the current settings, a patient is suspected positive, if they are experiencing 4 or more symptoms.
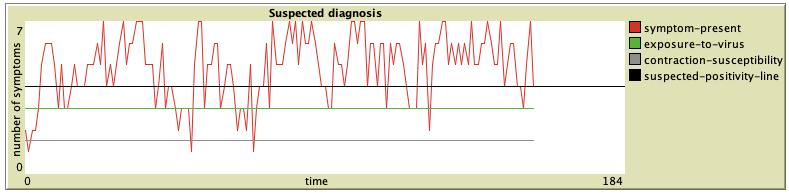
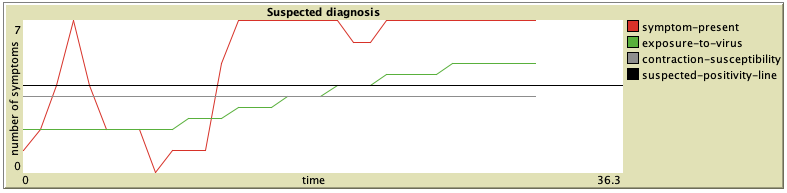
The plot shows the number of activated symptoms over time (per tick). As the exposure-to-virus slider is adjusted, the green line moves up or down accordingly. The same goes for contraction-susceptibility and the grey line. A person with very low contraction-susceptibility (e.g. a young, healthy, active person, or, quite newly, also a person that has been vaccinated) will be much less likely to cross the black suspected-positivity-line, even if the exposure-to-virus rate is very high. Similarly, an elderly person with long-term respiratory issues would be considered very contraction susceptible, and even a low exposure-to-virus rate might cause such person to cross the suspected-positivity-line.
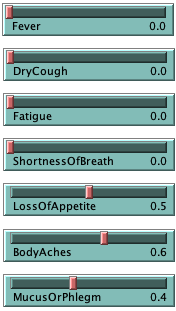
With the sliders per symptom on the right side of the model, it is possible to intervene on a specific symptom by curing it (by lowering it to 0.0). Similarly, it is possible to first set up the network with several symptoms (of identical or differing values) and see how such a patient progresses over time.
The probability of becoming activated for each symptom is represented as 1 / 1 + e^(a(b - A)).
Here, A represents the total amount of stress on a symptom (slider of symptom + exposure-to-virus + the activity of it's neighbors). In other words, A shows us how much pressure there is on the symptom to become activated.
a controls the steepness of the probability function and represents how much will a change in A affect the chance of activating the symptom. The parameter a has been adjusted according to the probability of a hospitalized COVID-19 patient developing the symptom.
And finally, b represents the threshold of a symptom - the higher b, the more activity in the neighbors such symptom needs to become activated. To take in consideration false positivity, the parameter b has been adjusted according to the combined probabilities of a patient developing said symptom while sick with the COVID-19 virus, the common flu, or cold. For example, for a symptom with the probability of development of 0.90199, the threshold has been set to 0.09801. The more common a symptom is, the higher are the chances of it appearing on it's own (not being caused/followed by other symptoms).
4. Results
To demonstrate the use of this simulation, 2 specifically preset runs will be used to illustrate.
A. First, we have a young man coming to the simulation clinic. He has a Fever (1.0), a DryCough (0.6) and is experiencing some ShortnessOfBreath (0.5). He has no other symptoms. He is otherwise healthy, active and a non-smoker, therefore his contraction-susceptibility has been set to 0.0. If we gradually increase his exposure-to-virus over about 30 ticks, we can watch other symptoms develop at a steady rate. Around 12 ticks the man has been first diagnosed as suspected positive by the model.
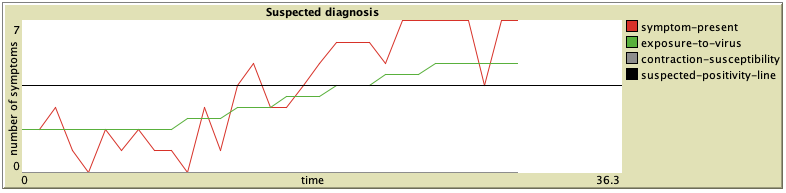
B. Second, an elderly lady arrives at the clinic. She has recently been experiencing LossOfAppetite (1.0) and her BodyAches (0.8). Otherwise, she has no other symptoms. Due to her age and a smoking habit, her contraction-susceptibility has been set to 0.7. Let us gradually increase her exposure-to-virus over about 30 ticks to compare with the man from case A:
We can see that not only the symptoms developed much faster (we were already able to first diagnose the lady at about 2 ticks), but the development has also been much steeper, causing the lady to experience almost all of the 7 main symptoms for the majority of the time. Hopefully, she got all better soon.
5. Conclusion
The main issue has, obviously, been data collection. As has been stated under Problem definition, the current sources are not only scarce, but also unreliable. However, I do believe that the positive side of my simulation is that it may be easily adjusted (symptoms added, thresholds edited, suspected-positivity-line moved, etc.) once we learn more about the disease. For now it may serve as a skeleton and, frankly, may be quite as reliable for the purposes of self-diagnosis as many of the suspiciously expensive home-testing kits available online.
Expanding the model using specific weights for the individual links between symptoms, if such are discovered during the ongoing COVID-19 research, could also bear very interesting fruits.
6. Code
Sources
WebMD. (2021). Signs of Coronavirus (COVID-19). [online] Available at: [1].
Health Service Executive, Ireland. (2021). Symptoms of coronavirus. [online] Available at: [2].
Who.int. (2019). 2019 Novel Coronavirus. [online] Available at: [3].
Health Infomation and Quality Authority, Ireland. (2020). Evidence summary for COVID-19 viral load over course of infection. [online] Available at: [4].